Molecular Dynamics
GROMACS is one of the most widely used open-source and free software codes in chemistry, used primarily for dynamical simulations of biomolecules.
GROMACS, developed by an international collaboration steered by KTH, is the major free and open source software package for biomolecular simulation. Its traditional strength has been its exceptionally high efficiency. Advances in hardware and software have enabled GROMACS simulations of macromolecules with hundreds of thousands of atoms, but the available simulation lengths are still orders of magnitude below important biological processes. To reduce the computational cost of calculating long-range interactions particle-mesh methods and 3D Fast Fourier Transform (FFT) are used to solve the Poisson solver in reciprocal space. Because all other interactions in molecular dynamics simulations are local, the 3D FFT becomes the bottleneck in parallelisation as it requires two global transposes of the charge grid over all the processors.
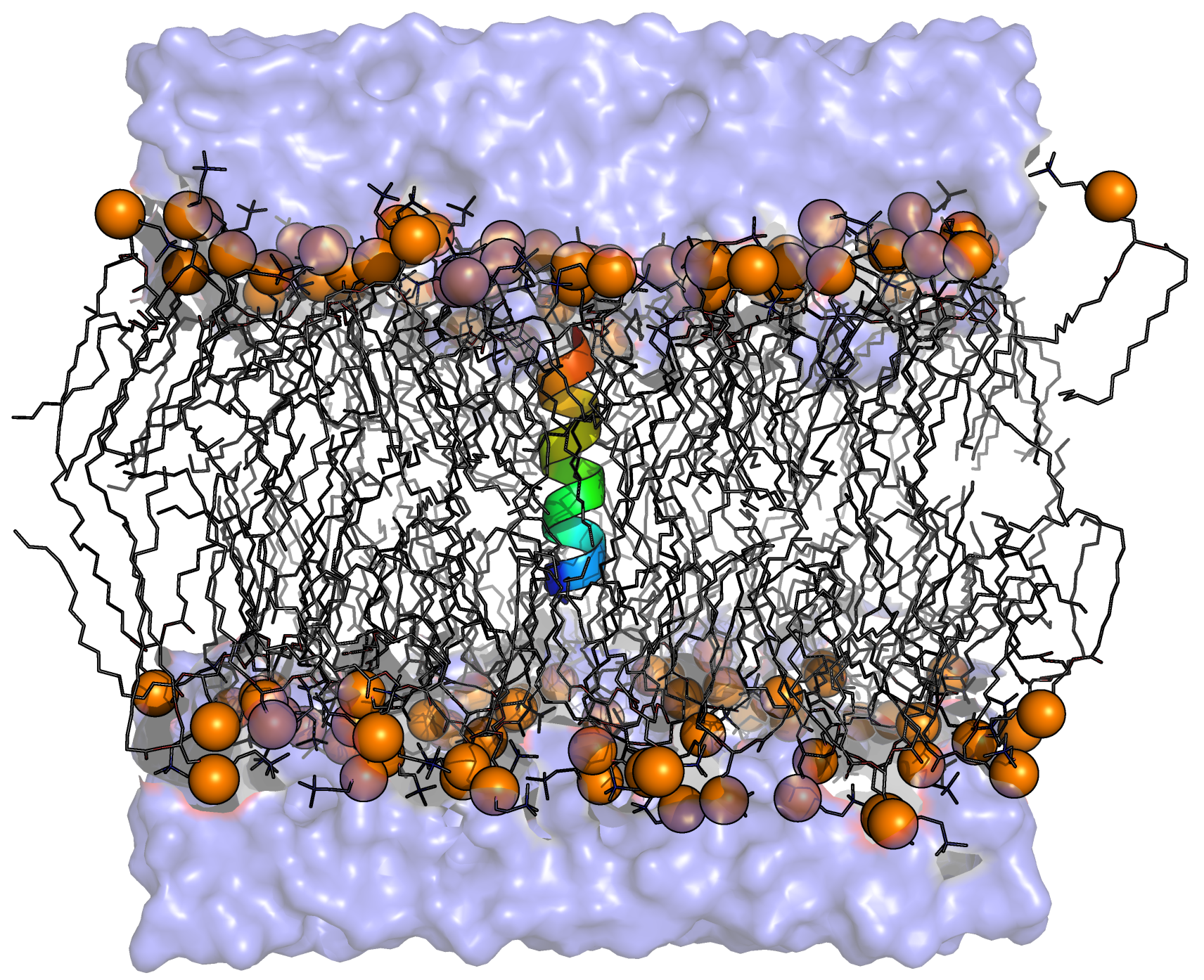
The objective for GROMACS in the DEEP-SEA consortium is to address the FFT performance bottleneck by designing and developing a GROMACS FFT library that is based on the DEEP-SEA software stack to exploit accelerators and novel hardware. In particular, we will demonstrate the usage of the DaCe framework to enhance the scalability of the FFT on a variety of different platforms, including distributed heterogeneous hardware. The success of this will be demonstrated by showing increased on-node performance of the GROMACS FFT solver and strong scalability to a large number of processes.